Who Are We?
Our work centers around multi-scale modeling in life sciences research and education. Located in the Biochemistry Department at the University of Nebraska-Lincoln, the focus of our efforts is three-fold:
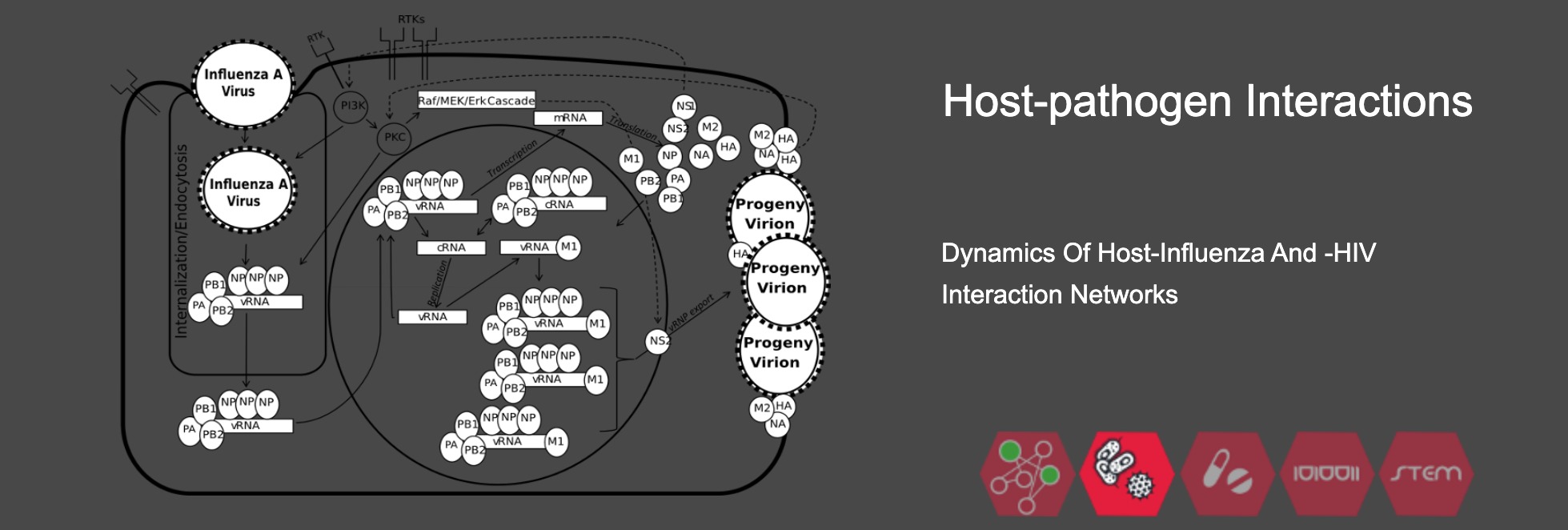
1) Better understand the dynamics of molecular and cellular mechanisms in complex networks under healthy as well as abnormal conditions. Examples of our work include the human immune system and plant root microbial interactions;
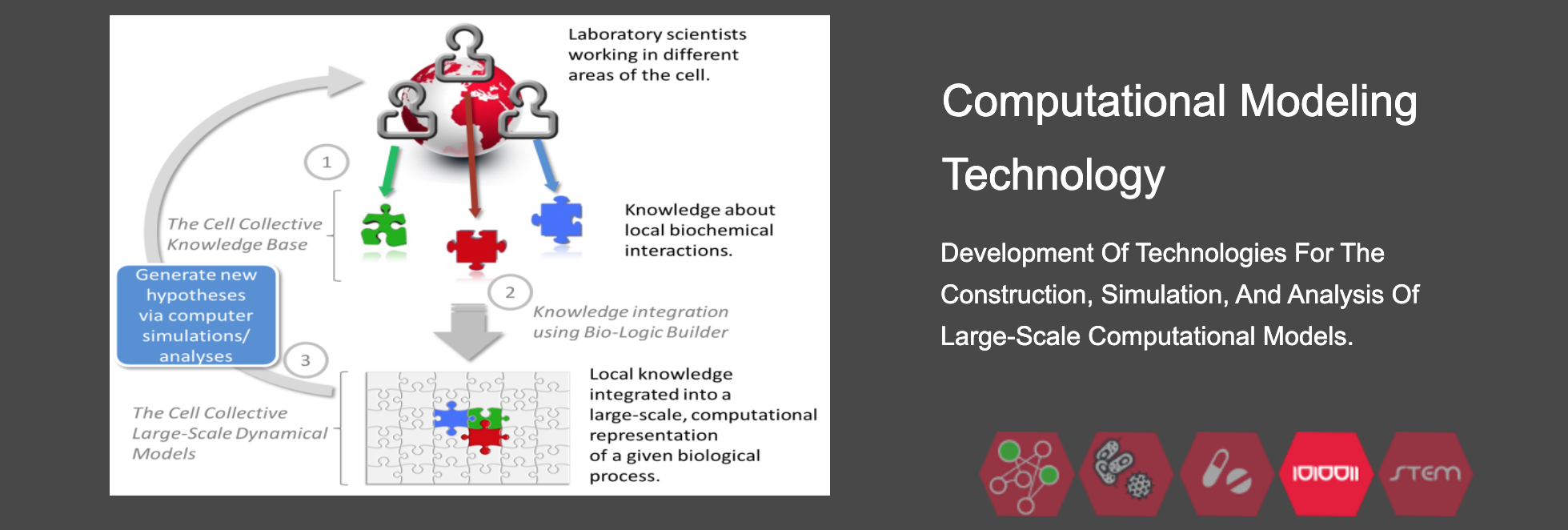
2) Make computational biology, modeling, data analysis and visualization broadly accessible to laboratory researchers, translational scientists, and professionals through easy-to-use cutting-edge software technologies, and
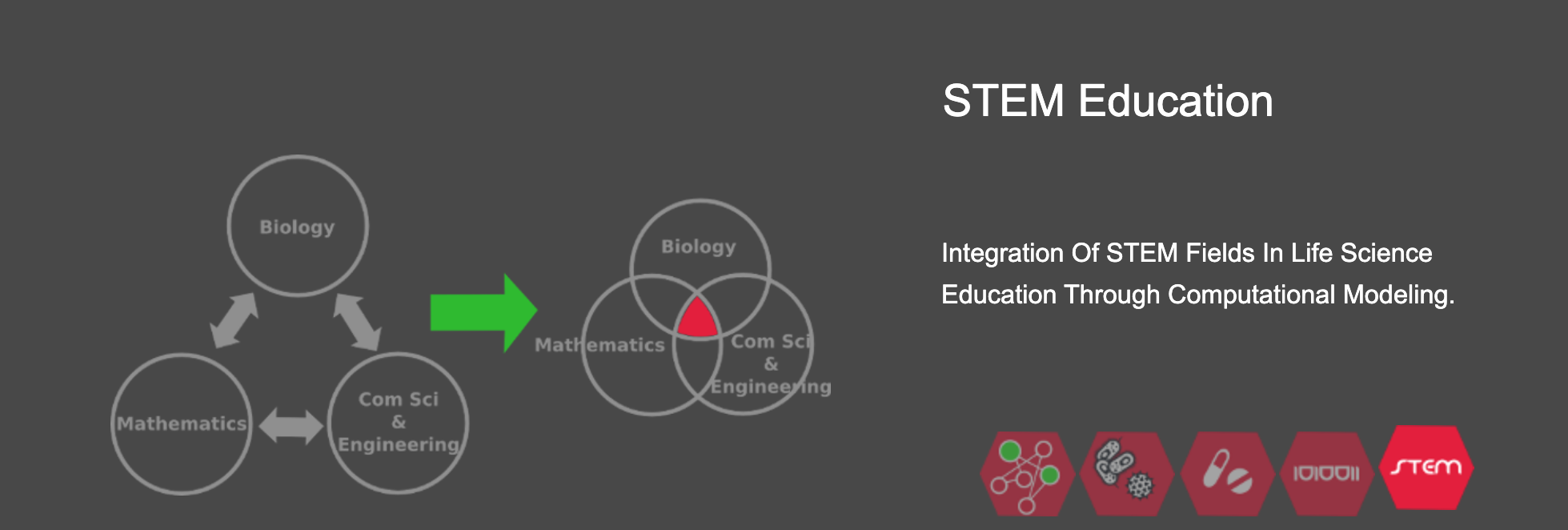
3) Enable computational modeling and simulation as an educational tool, whereby life sciences students can learn about the dynamic and complex nature of biological processes and diseases by building, breaking, simulating, and analyzing computational models of these biological systems. Our simulation-driven approach has now been adopted by tens of institutions in the US and worldwide, ranging from high schools to undergraduate and graduate-level life sciences courses.
Publications
Wertheim K.Y., Puniya B.L., La Fleur A., Shah, A. R., Barberis M., Helikar T*. 2021. Multi-Approach and Multi-Scale Model of CD4+ T Cells Predicts Switch-Like and Oscillatory Emergent Behaviors in Inflammatory Response to Infection. PLoS Computational Biology. 17 (8), e1009209
Yook J.S., You M., Kim J., Toney A. M., Fan R., Lal Puniya B., Helikar T., Vaulont S., Deschemin J. C., Okla M., Xie L., Manik G., Rauault T. A., Lee J., Chung, S. 2021. Coordination of beige adipogenesis with IRP, hepcidin, and HIF2α-dependent iron mobilization. Proceedings of the National Academy of Sciences. 118 (40)
Ostaszewski, M., Niarakis, A., Mazein, A., …. Helikar, T., Puniya, B., …. Schneider, R. 2021. COVID-19 Disease Map, a computational knowledge repository of SARS-CoV-2 virus-host interaction mechanisms. Molecular Systems Biology. 17:e10387
News
Contact Us
You can find us in the Biochemistry Department at the University of Nebraska-Lincoln, located in the Beadle Center.
University of Nebraska-Lincoln Beadle Center
1901 Vine St Lincoln, Nebraska 68503
email: thelikar2@unl.edu
tel: 402-472-3530